Computational Project#
Definition#
Project
A collection of artifacts (i.e., code, data, references, etc.) and metadata that provides a complete, traceable record of a research activity, study, or analysis.
Good Scientific Practice#
The question of what should, and what should not, be part of a project in computational science is determined by whatever is considered good scientific practice.
A detailed list of recommendations of what are good practices for a computational science project is provided by Wilson et al. (2017)1Wilson, G., et al. (2017). Good enough practices in scientific computing. PLoS Comp Bio. https://doi.org/10.1371/journal.pcbi.1005510.. Provided here-below is a condensed representation:
Preserve: Save and back up original raw data in multiple locations.
Structure: Create analysis-ready, tidy data with unique identifiers.
Traceability: Document all processing steps and archive in a DOI-issuing repository.
Clean Code: Use modular functions, meaningful names, and avoid duplication.
Dependencies: Use/test external libraries and make requirements explicit.
Documentation: Provide header comments, a test dataset, and a DOI for the code.
Roadmap: Maintain a project overview and shared to-do list.
Sync: Define communication strategies and explicit licensing.
Citation: Ensure the project is citable for others.
Version Control: Use a VCS (like Git) for all human-created files.
Frequency: Share small, frequent changes rather than large, rare ones.
Logs: Maintain a
CHANGELOGand mirror work to a remote server.
Format: Use plain text formats (like Markdown) to allow for version control.
Tools: Use platforms with rich formatting and automated reference management.
2026 Update#
Almost 10 years after Wilson et al. (2017) an appendix is expedient:
Disclosure: State where AI (e.g., LLMs) was used for artifact (code, logic, text) generation and modification.
Verification: Audit all AI-generated artifacts for errors.
Prompts & Instructions: Archive prompts used to generate critical artifacts, like code or analysis workflows.
Isolation: Work with dedicated virtual environments to enforce dependency control.
Pinned Dependencies: Use lockfiles to ensure exact version parity.
Virtualization: Containerize workflows to encapsulate the software environment.
Specs: Document GPU model, VRAM, and driver versions (e.g., CUDA).
Virtualization: Use VMs to get a full documentation of the hardware stack used.
Unit Tests: Write tests for functions to ensure logic stays consistent.
Fixed Seeds: Set and record seeds for all random number generators.
Separation of Concerns (SoC)#
When organizing scientific projects (especially in data science) one principle stands above the rest:
The separation of logic, parameters, environment, and data
This principle, known as Separation of Concerns (SoC), is fundamental for reproducibility, clarity, and maintainability. It dictates how the digital artifacts of a project should be physically organized on disk, ensuring that a change in one domain (e.g., running the code on a different computer) does not require a change in another (e.g., modifying the core logic).
At the most basic level, Separation of Concerns implies creating distinct subdirectories to separate the code from its configuration and the data it operates on:
my_data_science_project/
│
├── config/ # Directory for configuration files
├── data/ # Directory for input data
└── code/ # Directory for programming logic
.
.
However, a robust project structure goes deeper. Best practice requires explicitly separating logic, parameters, environment/state, and the data inputs/outputs. Hardcoding data paths or attempting to store massive datasets within a repository is highly discouraged. By introducing an .env file and distinct input/output directories, the project becomes modular and environment-agnostic:
my_data_science_project/
│
├── config/ # Parameters: Centralized parameterizations (YAML/TOML)
├── .env # Environment: Local paths and secrets (DO NOT COMMIT)
│
├── scripts/ # Logic: Python scripts for exploration and analysis
│
├── data/ # Data (Input): Kept small or used for symlinks
└── results/ # Data (Output): Plots, figures, tables
.
.
Using Environment Variables
The .env file defines local variables, such as absolute paths to external storage drives where the actual training data or output directories reside. These paths can be loaded dynamically using the python-dotenv package:
import os
from dotenv import load_dotenv
# Load variables from the local .env file
load_dotenv()
# Safely fetch environment-specific paths
output_dir = os.getenv("OUTPUT_DIR")
Best Practice: Securing .env files
The .env file must never be committed to version control, as it may contain sensitive paths or API keys. It should be explicitly added to the .gitignore file. Instead, an .env.example file should be committed, providing a blank template so collaborators know which variables must be defined on their respective systems.
As complexity grows, the advanced case requires splitting these concerns even further. Logic is divided into executable routines (scripts/) and reusable modules (src/). Data is divided into discrete stages (raw/, interim/, final/) to trace lineage and document processing steps:
my_data_science_project/
│
├── data/ # Data stages
│ ├── raw/ # Original, immutable data dumps
│ ├── interim/ # Intermediary data from filtering/cleaning
│ └── final/ # Cleaned, tidy data used for analysis
│
├── src/ # Reusable logic: The core Python package
├── scripts/ # Executable logic: Batch execution scripts
│
├── config/ # Parameters
├── .env # Environment state
└── results/ # Output deliverables
.
.
Finally, to make the project entirely self-describing, essential metadata and documentation are added to the root structure:
my_data_science_project/
│
├── data/ # Data stages
├── src/ # Reusable logic
├── scripts/ # Executable logic
├── results/ # Outputs
├── config/ # Parameters
│
├── docs/ # Documentation source files (e.g., Sphinx, MkDocs)
├── pyproject.toml # Project metadata and dependencies (SSOT)
├── README.md # Project overview and instructions
├── LICENSE # License governing usage
├── .env.example # Template for the .env file (committed to version control)
└── .gitignore # Files and directories to ignore in Git
Quality and Control#
Once the core concerns are separated, an extensive structure integrates dedicated components for quality assurance and automation. Testing and benchmarking are critical for ensuring that the logic performs accurately and efficiently over time.
Furthermore, modern scientific computing relies heavily on Continuous Integration and Continuous Deployment (CI/CD). By including configuration files for platforms like GitHub or GitLab, the execution of tests, benchmarks, and documentation builds can be fully automated whenever changes are made to the repository.
my_data_science_project/
│
├── tests/ # Quality Control: Automated unit and integration tests
│ ├── requirements.txt # Dependencies specifically for testing (e.g., pytest)
│ └── test_cleaning.py # Test suites corresponding to src modules
│
├── benchmark/ # Quality Control: Performance and scaling tests
│ └── run_benchmarks.py # Scripts to track execution time and resource usage
│
├── .github/ # Automation: GitHub-specific configurations
│ └── workflows/ # GitHub Actions CI/CD pipelines (YAML)
│
├── .gitlab-ci.yml # Automation: GitLab CI/CD pipeline configuration
.
.
Note on CI/CD: A project typically utilizes either
.github/workflows/(if hosted on GitHub) or.gitlab-ci.yml(if hosted on GitLab), but rarely both simultaneously. They are included together here simply for illustrative purposes.
4 Pillars of SoC#
The Four Pillars of SoC
True computational reproducibility mandates the rigid Separation of Concerns (SoC) across four distinct, isolated pillars. Failure to isolate these concerns leads to security vulnerabilities (credentials in Git), rigid codebases, non-reproducible environments, and data integrity failures.
Effective data handling is critical for computational efficiency, reproducibility, and scientific integrity. Within a modern computational project, data must not be treated as an ephemeral byproduct but as a first-class citizen governed by a deliberate strategy.
This strategy hinges on the strict application of the Separation of Concerns (SoC) principle. Concerns are separated into four distinct pillars, which only intersect at the moment of execution within a controlled script:
Environment (Software stack and runtime context)
Configuration (Static algorithmic parameters)
Code (Reusable logic vs. execution coordination)
Data (Actual data vs. state declarations)
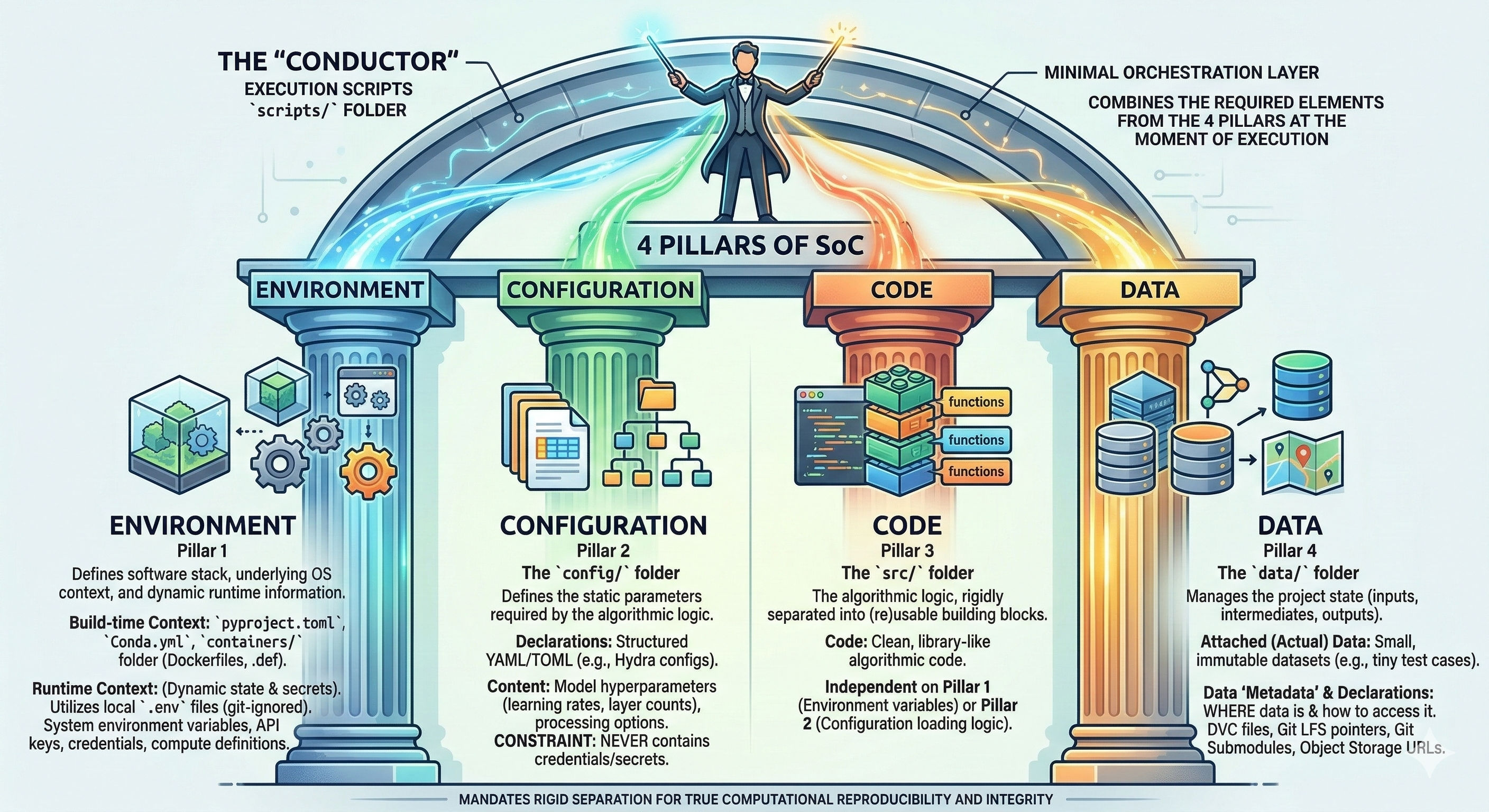
The Four Pillars of SoC#
The “conductor”
Execution scripts (under scripts/) act as minimal orchestration layer that combines the required elements from the 4 pillars:
Pillar 1
Defines the software stack, underlying OS context, and dynamic runtime information.
Build-time Context: Defines the static software image.
Declarations:
pyproject.toml,Conda.yml.Definition Artifacts:
containers/folder holding.deforDockerfiles.
Runtime Context: Defines the dynamic execution state and secrets.
Mechanisms: Utilizes local
.envfiles (git-ignored).Content: Passed via system environment variables; contains credentials, dynamic API keys, and compute resource definitions.
Pillar 2
The config/ folder
Defines the static parameters required by the algorithmic logic.
Declarations: Structured YAML/TOML files (e.g., Hydra configs).
Content: Model hyperparameters (learning rates, layer counts), data processing options, and application-specific settings.
Constraint: Never contains credentials or secrets (passed via runtime environment).
Pillar 3
The src/ folder
The algorithmic logic, rigidly separated into (re)usable building blocks.
Code: Clean, library-like algorithmic code.
Independent on Pillar 1 (Environment variables) or Pillar 2 (Configuration loading logic).
Pillar 4
The data/ folder
Manages the project state (inputs, intermediates, outputs). A strict distinction is made between attached data and data definitions (“metadata”).
Attached (Actual) Data:
Should only contain small, immutable datasets localized to the repository (e.g., tiny test cases).Data “Metadata” & Declarations:
Defines WHERE data is and how to access it, rather than hosting it.Declarations: Contains DVC (Data Version Control) files, Git LFS pointer files, or Git Submodules.
Access Info: Object Storage URLs, bucket names, and endpoint paths.
Project Scaffolding#
A fundamental step toward following best practices in any data or software project is to start with a well-structured layout. This presents a consistent project architecture that is applicable to wide range of projects, from simple analytical pipelines to robust software packages.
Naturally, the exact specifics will shift depending on the subject matter, the technologies used, and the overall purpose of the work. However, there is a golden rule to maintain sanity and collaboration: the root of a repository should always look the same.
Adhering to a standardized root directory structure reduces friction. It facilitates a faster setup, makes the codebase instantly recognizable to collaborators, and allows anyone to quickly orient themselves without getting lost in a maze of arbitrarily named folders.
We suggest adopting the following structure for the root folder, commonly referred to as the Project Scaffolding.
- 📄
- 📄
- 📄
- 📄
- 📄
- 📄
- 📄
- 📄
- 📄
- 📄
- 📄
-
- 📄
- 📂
- 📂
- 📂
-
- 📂
-
- 📂
- 📂
- 📂
-
- 📂
- 📂
-
- 📂
- 📂
- 📄
- 📂
- 📂
- 📂
-
- 📂
🏠 Project Root
The root folder of a project should only contain generic files and foldernames.
Select a file or folder to read more about its purpose and content
Click on the 📁 signs to expand a folder.
A 🔸 indicates a specific context in which the element is relevant.
📖 README.md
The front page of a repository.
It should serve as nexus for all metadata related to a project.
Especially for bigger projects this also means that the `README.md` might **not** contain all metadata directly, but links and instructions on how to find all metadata.
📦 pyproject.toml
The modern standard for defining build systems, dependencies, and tool settings (like ruff, pytest, or black).
📄 .env.example
A safe template for the .env file. Unlike the actual .env file, this template must be committed to version control.
It provides empty or safe default values to demonstrate exactly which environment variables a collaborator needs to define on their own system to execute the project successfully.
A minimal example of its contents:
# Define the absolute path to the local data directory
RAW_DATA_DIR=/absolute/path/to/local/data/
# Provide the API key for external service access
EXTERNAL_API_KEY=<insert_api_key_here>
🙈 .gitignore
Specifies intentionally untracked files that Git should ignore (e.g., .env, __pycache__, local data, build artifacts).
🔐 .env
Local environment variables and secrets (such as API keys, database passwords, and personal access tokens). This file should never be added to version control. It should be strictly ignored by Git to prevent leaking sensitive security credentials.
We recommend to add the .env file to .gitignore and provide a .env.example file along with a repository so to document how exactly the .env file should be structured.
⚙️ .gitlab-ci.yaml
The CI/CD pipeline definition for GitLab. It declares stages, jobs, and rules that automatically run tests, build artifacts, and deploy your project whenever changes are pushed to the repository.
A single YAML file at the repository root controls the entire automation pipeline. Each job specifies a Docker image, a script to execute, and the conditions under which it should run. GitLab Runner picks up these jobs and executes them in isolated containers.
⚙️ .github/
The hidden configuration directory for GitHub-specific features: CI/CD workflows, issue templates, pull request guidelines, and more.
⚙️ workflows/
GitHub Actions workflow files (YAML) that define automated CI/CD pipelines. Each file describes a workflow triggered by events such as pushes, pull requests, or scheduled runs to test, build, and deploy your project.
Unlike GitLab's single-file approach, GitHub Actions uses one YAML file per workflow inside this directory, making it straightforward to maintain separate pipelines for testing, releasing, and documentation deployment.
🤖 AI_USAGE.md
Transparency document detailing how LLMs were used in the creation of code or documentation.
⚖️ LICENSE
The legal framework for a project. It explicitly defines how others can use, modify, and distribute the code and data. Even for internal or proprietary projects, having a clear license is essential.
📝 CITATION.cff
Plain text YAML file specifying how this project should be cited.
This should be a plain text file with machine-readable citation information.
A minimalistic example of what this file looks like:
cff-version: 1.2.0
message: "If you use this software, please cite it using these metadata."
authors:
- family-names: "Lisa"
given-names: "Mona"
orcid: "https://orcid.org/xxxx-xxxx-xxxx-xxxx
title: "My Awesome Research"
doi:
For more details visit the cff-format documentation.
🤝 CONTRIBUTING.md
The onboarding guide for new developers. It outlines the step-by-step process for submitting bug reports, requesting features, and creating pull requests (PRs) that adhere to the project's standards.
🛡️ CODE_OF_CONDUCT.md
Establishes the community standards and expected behavior for everyone interacting with the project. It helps ensure a welcoming, inclusive, and professional environment for all contributors.
📊 Data
The root data directory. We use a strict data pipeline separating raw inputs from processed outputs. Expand the folder and select a subfolder on the left to learn more about its specific rules.
📖 README.md
Information specific to the data related to this project.
📁 raw/
Immutable, original data. Do not edit these files. (Note: This also includes pointers/URL links to large external datasets that cannot be stored directly in Git.)
📁 interim/
Intermediate data that has been cleaned or transformed, but is not yet ready for final analysis.
📁 final/
Final, processed datasets ready for modeling, publication, or deployment.
🚀 Scripts
Executable scripts, job launchers, or data preprocessing entry points. These should generally be runnable from the command line.
📁 drafts/
A scratchpad folder for experimental scripts that are not yet production-ready or integrated into the main workflow.
📓 Notebooks
Jupyter notebooks for exploratory data analysis (EDA), prototyping, and interactive visualization.
📁 drafts/
A scratchpad folder for experimental notebooks that are not yet production-ready or integrated into the main workflow.
📦 Containers
Centralized location for all container definitions. Your *.def and Dockerfile files are here.
⚙️ Configuration
Centralized YAML/TOML files for parameters. This allows changing experiments without modifying code.
💻 Source Code (src/)
Location for all reusable and installable code.
📁 mypkg/
A particular package containing functions, and business logic.
📈 results/
A dedicated directory for the final outputs of of a project. If the project involves data analysis, research, or machine learning, this folder holds the generated artifacts such as figures, summary tables, trained models, and compiled reports.
🧪 Testing Suite
Ensures code reliability via automated testing. A well-structured test suite separates individual component tests from full workflow tests.
📁 unit/
Directory for unit tests. These tests are designed to verify that individual functions, classes, or methods operate correctly in strict isolation.
📁 integration/
Directory for integration tests. These tests verify that multiple modules, databases, or external services function together properly as a unified system.
📄 test_mypkg.py
A standard Python test file (often utilizing pytest). This file contains specific test cases and assertions designed to validate the functionality of the associated mypkg module.
💡 Examples
Minimal, end-to-end usage demos to help new users get started quickly.
📖 Documentation
Contains extended documentation specific content, like an online documentation (e.g., with Sphinx or MKDocs).
⏱️ Benchmarks
Performance tracking scripts to measure execution time, memory usage, and scaling.
Template Repository#
Because the root structure and core configuration files of most projects share a highly consistent layout, the creation of a standardized project template is highly recommended. Utilizing a template eliminates the repetitive process of manually recreating scaffolding, ensuring that every new endeavor begins with best practices already in place.
By defining a baseline architecture — complete with placeholder directories, standard configuration files (such as pyproject.toml or .gitignore), and foundational documentation — consistency is maintained across multiple projects.
This approach drastically reduces initial setup time, prevents structural drift between different analyses, and ensures that critical components like licensing and dependency management are never overlooked.
A pre-configured template repository is provided below. It is designed to serve as a comprehensive, ready-to-use foundation for any Python-based project:
Templates
A good starting point for finding project templates for various purposes is cookiecutter.io.
A project template following closely the structure presented is available on GitHub: T4D’s Python Project Template.
Delimination#
A well-planned project structure naturally encourages consolidating data, scripts, tests, and documentation into a single repository. The primary advantage of this “monolithic” approach is simplicity: all elements of a project can be tracked, versioned, and shared as one cohesive unit.
However, consolidating absolutely everything into a single repository can become problematic as a project scales. Issues may eventually be encountered regarding:
Distribution & Licensing:
Different components often need to be shared under different restrictions. For instance, custom code might be open-source, but it may rely on a proprietary third-party dataset that explicitly prohibits public redistribution.Complexity & Bloat:
Massive repositories can quickly become unwieldy, obscuring the relationship between individual elements, slowing down Git operations, and making the commit history difficult to navigate.
To avoid a bloated monolithic repository while maintaining a unified workflow, multiple smaller, focused repositories can be linked together using git submodules.
This allows a main project to include an external element (such as a raw dataset or a shared utility library) by securely pointing to a specific commit in another Git repository.
Deciding exactly when to break a project into multiple repositories is ultimately somewhat arbitrary, but a separate repository is generally recommended if:
An element (e.g., a curated dataset) is broadly useful in isolation for entirely different projects.
An element (e.g., public-facing documentation or a web dashboard) needs to evolve or be deployed independently of the main analytical code.
An element is subject to different ownership, access rights, or terms of use.
📝 A Note on Manuscripts
For papers, book chapters, or theses, the creation of a dedicated, separate repository is strongly recommended.
Resources from a primary analysis repository (such as auto-generated figures or tables) can still be easily referenced by including the analysis repository as a submodule.
However, a written manuscript and the underlying code analysis follow fundamentally different editing lifecycles; their separation keeps the writing history clean and prevents the codebase from being flooded with minor typo fixes.
Sources:
https://packaging.python.org/en/latest/guides/writing-pyproject-toml/
https://peps.python.org/
https://setuptools.pypa.io/en/latest
https://docs.python.org/3.12/tutorial/modules.html
Wilson, G., Bryan, J., Cranston, K., Kitzes, J., Nederbragt, L., and Teal, T. K. (2017). Good enough practices in scientific computing. PLoS Computational Biology, 13(6), e1005510. https://doi.org/10.1371/journal.pcbi.1005510
Slides “STA472 Good Statistical Practice”, Week 2 - Reproducibility and data, Project organization and software (by Eva Furrer)
https://setuptools.pypa.io/en/latest
https://docs.python.org/3.12/tutorial/modules.html
Marwick, B., Boettiger, C., and Mullen, L. (2018). Packaging Data Analytical Work Reproducibly Using R (and Friends). The American Statistician, 72(1), 80–88. https://doi.org/10.1080/00031305.2017.1375986